Pseudomonas aeruginosa is a bacterium responsible for severe nosocomial infections, life-threatening infections in immunocompromised persons, and chronic infections in cystic fibrosis patients. The bacterium's virulence depends on a large number of cell-associated and extracellular factors. Cell-to-cell signaling systems control the expression and allow a coordinated, cell-density–dependent production of many extracellular virulence factors. We discuss the possible role of cell-to-cell signaling in the pathogenesis of P. aeruginosa infections and present a rationale for targeting cell-to-cell signaling systems in the development of new therapeutic approaches.
Pseudomonas aeruginosa as a Human Pathogen
Pseudomonas aeruginosa, an increasingly prevalent opportunistic human pathogen, is the most common gram-negative bacterium found in nosocomial infections. P. aeruginosa is responsible for 16% of nosocomial pneumonia cases, 12% of hospital-acquired urinary tract infections, 8% of surgical wound infection, and 10% of bloodstream infections. Immunocompromised patients, such as neutropenic cancer and bone marrow transplant patients, are particularly susceptible to opportunistic infections. In this group of patients, P. aeruginosa is responsible for pneumonia and septicemia with attributable deaths reaching 30%. P. aeruginosa is also one of the most common and lethal pathogens responsible for ventilator-associated pneumonia in intubated patients, with directly attributable death rates reaching 38%. In burn patients, P. aeruginosa bacteremia has declined as a result of better wound treatment and dietary changes (removal of raw vegetables, which can be contaminated with P. aeruginosa, from the diet). However, P. aeruginosa outbreaks in burn units are still associated with high (60%) death rates. In the expanding AIDS population, P. aeruginosa bacteremia is associated with 50% of deaths. Cystic fibrosis (CF) patients are characteristically susceptible to chronic infection by P. aeruginosa, which is responsible for high rates of illness and death in this population. The capacity of P. aeruginosa to produce such diverse, often overwhelming infections is due to an arsenal of virulence factors. Many extracellular virulence factors secreted by P. aeruginosa have been shown to be controlled by a complex regulatory circuit involving cell-to-cell signaling systems that allow the bacteria to produce these factors in a coordinated, cell-density–dependent manner. In this article we describe major virulence factors of P. aeruginosa and the possible involvement of cell-to-cell signaling in the pathogenesis of acute P. aeruginosa infection. We also summarize data suggesting that these regulatory systems could be exploited for the design of therapeutic interventions.
Pathogenesis of P. aeruginosa Infections
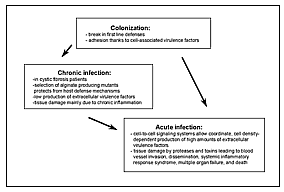
P. aeruginosa (family Pseudomonadaceae), an aerobic, motile, gram-negative rod able to grow and survive in almost any environment, lives primarily in water, soil, and vegetation. However, despite abundant opportunities for spread, P. aeruginosa rarely causes community-acquired infections in immunocompetent patients. As a result, the pathogen is viewed as opportunistic. The different phases of P. aeruginosa infection are shown in Figure 2.
Colonization: The Predominant Role of Cell-Associated Virulence Factors
To initiate infection, P. aeruginosa usually requires a substantial break in first-line defenses. Such a break can result from breach or bypass of normal cutaneous or mucosal barriers (e.g., trauma, surgery, serious burns, or indwelling devices), disruption of the protective balance of normal mucosal flora by broad-spectrum antibiotics, or alteration of the immunologic defense mechanisms (e.g., in chemotherapy-induced neutropenia, mucosal clearance defects from cystic fibrosis, AIDS, and diabetes mellitus).
The first step in P. aeruginosa infections is colonization of altered epithelium. The pathogen colonizes the oropharynx of up to 6% and is recovered from the feces of 3% to 24% of healthy persons. In contrast, up to 50% of hospitalized patients are at high risk for P. aeruginosa colonization. Adherence of P. aeruginosa to epithelium is probably mediated by type 4 pili similar to those of Neisseria gonorrhoeae. Several other nonpilus adhesins responsible for the binding to mucin have been described, but their role in the infection process remains unclear. Flagella, which are primarily responsible for motility, may also act as adhesins to epithelial cells.
From Colonization to Acute Infection: The Role of Extracellular Virulence Factors
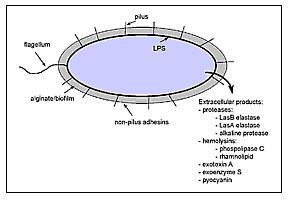
P. aeruginosa produces several extracellular products that after colonization can cause extensive tissue damage, bloodstream invasion, and dissemination (Figure 1). In vivo studies have shown that mutants defective in the production of exotoxin A, exoenzyme S, elastase, or alkaline protease are essential for maximum virulence of P. aeruginosa; however, the relative contribution of a given factor may vary with the type of infection. Many of these factors are controlled by regulatory systems involving cell-to-cell signaling. We will summarize the known biologic effects of the most-studied extracellular virulence factors associated with acute P. aeruginosa infection.
Exotoxin A is produced by most P. aeruginosa strains that cause clinical infections. Like diphtheria toxin, P. aeruginosa exotoxin A catalyzes ADP-ribosylation and inactivation of elongation factor 2, leading to inhibition of protein biosynthesis and cell death. Exotoxin A is responsible for local tissue damage, bacterial invasion, and (possibly) immunosuppression (19). Purified exotoxin A is highly lethal for mice which supports its role as a major systemic virulence factor of P. aeruginosa.
Exoenzyme S is also an ADP-ribosyl transferase, but unlike exotoxin A, it preferentially ribosylates GTP-binding proteins such as Ras. This exoproduct is responsible for direct tissue destruction in lung infection and may be important for bacterial dissemination.
Two hemolysins, phospholipase C and rhamnolipid, produced by P. aeruginosa, may act synergistically to break down lipids and lecithin. Both may contribute to tissue invasion by their cytotoxic effects. Rhamnolipid, a rhamnose-containing glycolipid biosurfactant, has a detergentlike structure and is believed to solubilize the phospholipids of lung surfactant, making them more accessible to cleavage by phospholipase C. The resulting loss of lung surfactant may be responsible for the atelectasis associated with chronic and acute P. aeruginosa lung infection. Rhamnolipid also inhibits the mucociliary transport and ciliary function of human respiratory epithelium. However, the relative role of rhamnolipid in acute or chronic infection is not known.
Proteases are assumed to play a major role during acute P. aeruginosa infection. P. aeruginosa produces several proteases including LasB elastase, LasA elastase, and alkaline protease. The role of alkaline protease in tissue invasion and systemic infections is unclear; however, its role in corneal infections may be substantial. The ability of P. aeruginosa to destroy the protein elastin is a major virulence determinant during acute infection. Elastin is a major part of human lung tissue and is responsible for lung expansion and contraction. Moreover, elastin is an important component of blood vessels, which rely on it for their resilience. The concerted activity of two enzymes, LasB elastase and LasA elastase, is responsible for elastolytic activity. Elastolytic activity is believed to destroy elastin-containing human lung tissue and cause the pulmonary hemorrhages of invasive P. aeruginosa infections. LasB elastase is a zinc metalloprotease that acts on a number of proteins including elastin. LasB elastase is highly efficient, with a proteolytic activity approximately 10 times that of P. aeruginosa alkaline protease and an activity toward casein approximately four times that of trypsin. The LasA elastase is a serine protease that acts synergistically with LasB elastase to degrade elastin. LasA elastase nicks elastin, rendering it sensitive to degradation by other proteases such as LasB elastase, alkaline protease, and neutrophil elastase. Both LasB elastase and LasA elastase have been found in the sputum of CF patients during pulmonary exacerbation. However, the role of LasB elastase in tissue destruction during the chronic phase of CF is less clear. It has been postulated that during this phase, antibodies present in high titers neutralize LasB elastase, and elastin damaged by minute amounts of LasA is degraded mostly by neutrophil elastase. LasB elastase degrades not only elastin but also fibrin and collagen. It can inactivate substances such as human immunoglobulins G and A, airway lysozyme, complement components, and substances involved in protecting the respiratory tract against proteases such as a-1-proteinase inhibitor and bronchial mucus proteinase inhibitor. Therefore, LasB elastase not only destroys tissue components but also interferes with host defense mechanisms. Studies in animal models show that mutants defective in LasB elastase production are less virulent than their parent strains, which supports the role of LasB elastase as a virulence factor.
Cell-to-Cell Signaling: A Global Regulation System of P. aeruginosa Extracellular Virulence Factors
Cell-to-Cell Signaling Systems
P. aeruginosa appears to control the production of many of its extracellular virulence factors by a mechanism that monitors bacterial cell density and allows communication between bacteria by cell-to-cell signaling. Bacteria are able to sense their environment, process information, and react appropriately; however, their ability to sense their own cell density, to communicate with each other, and to behave as a population instead of individual cells has only recently been understood. This phenomenon, called quorum-sensing or cell-to-cell signaling, is a generic phenomenon described in many gram-negative and gram-positive bacteria.
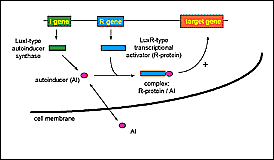
The first cell-to-cell signaling system described is the lux system, responsible for the cell-density–dependent control of bioluminescence genes by the marine bacteria Vibrio fischeri. Cell-to-cell signaling systems of gram-positive bacteria involve peptide pheromones as signals. In contrast, cell-to-cell signaling systems of gram-negative bacteria, with the exception of Ralstonia solanacearum, in which an endogenous fatty acid derivative has been suggested as a cell-to-cell signal, are composed of a small molecule called an autoinducer, which is synthesized by a LuxI-type autoinducer synthase and a LuxR-type transcriptional activator protein (R-protein) (Figure 3). The various autoinducers described in gram-negative bacteria are homoserine lactone-based molecules that differ between one another in length and substitutions on their acyl side chains. At low cell density, autoinducer is synthesized at basal levels and is thought to diffuse into the surrounding media, where it becomes diluted. With increasing cell density, the intracellular concentration of autoinducer increases until it reaches a threshold concentration. At this critical concentration, the autoinducer has been proposed to bind to a specific R-protein. The R-protein itself is not active without the corresponding autoinducer, and it is the R-protein/autoinducer complex that is proposed to bind to specific DNA sequences upstream of target genes enhancing their transcription. The resulting increase in expression of these genes can reach 1,000-fold. The autoinducer, therefore, allows the bacteria to communicate with each other (cell-to-cell signaling), to sense their own density (quorum-sensing), and together with a transcriptional activator to express specific genes as a population instead of individual cells.
The las Cell-to-Cell Signaling System of P. aeruginosa
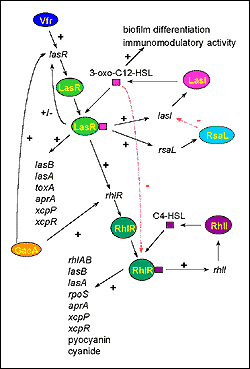
The first cell-to-cell signaling system described in P. aeruginosa was shown to regulate expression of LasB elastase and was therefore named the las system. The las cell-to-cell signaling system is composed of lasI, the autoinducer synthase gene responsible for the synthesis of 3-oxo-C12-HSL (N-[3-oxododecanoyl]-L-homoserine lactone, previously named PAI-1 or OdDHL), and the lasR gene that codes for a transcriptional activator protein (Figure 4). The las cell-to-cell signaling system regulates lasB expression and is required for optimal production of other extracellular virulence factors such as LasA protease and exotoxin A. LasI is the most sensitive gene to activation by LasR/3-oxo-C12-HSL. The preference for the lasI promoter allows an initial rapid rise in autoinducer synthesis, which increases the amount of 3-oxo-C12-HSL available to bind to LasR. This autoinduction hierarchy is responsible for a dramatic increase of expression of virulence genes (such as lasB) once a critical cell density has been reached. Recently, the las system has also been shown to activate the xcpP and xcpR genes that encode proteins of the P. aeruginosa secretory pathway. 3-oxo-C12-HSL alone has been suggested to contribute to the virulence of P. aeruginosa because it has some immunomodulatory activity . The las cell-to-cell signaling system is positively controlled by GacA, as well as by Vfr, which is required for the transcription of lasR. An inhibitor, RsaL, that represses the transcription of lasI, has also been described (de Kievit et al., unpub. data). The multiple regulatory levels of the las cell-to-cell signaling system and the various genes under its control highlight the importance of this system for P. aeruginosa.
The rhl Cell-to-Cell Signaling System of P. aeruginosa
P. aeruginosa has a second cell-to-cell signaling system, named the rhl system because of its ability to control the production of rhamnolipid. This system is composed of rhlI, the C4-HSL (N-butyrylhomoserine lactone, previously named PAI-2 or BHL) autoinducer synthase gene, and the rhlR gene encoding a transcriptional activator protein (Figure 4). This system regulates the expression of the rhlAB operon that encodes a rhamnosyltransferase required for rhamnolipid production. The rhl system is also necessary for optimal production of LasB elastase, LasA protease, pyocyanin, cyanide, and alkaline protease. Therefore, like the las cell-to-cell signaling system, the rhl system, sometimes referred to as vsm (virulence secondary metabolites), regulates the expression of various extracellular virulence factors of P. aeruginosa. Interestingly, the rhl system also regulates the expression of rpoS, which encodes a stationary sigma factor (dS) involved in the regulation of various stress-response genes.
The Cell-to-Cell Signaling Hierarchy in P. aeruginosa
Recent data have shown that the las and rhl cell-to-cell signaling systems of P. aeruginosa interact. Both systems are highly specific in that their respective autoinducers are unable to activate the transcriptional activator protein of the other system (i.e., 3-oxo-C12-HSL cannot activate RhlR, and C4-HSL cannot activate LasR). It has also been shown that the R-protein/autoinducer complexes prefer certain promoters that they will activate; LasR/3-oxo-C12-HSL preferentially activates lasB over rhlA, and RhlR/C4-HSL preferentially activates rhlA over lasB. However, neither system is completely independent of the other. The LasR/3-oxo-C12-HSL complex activates the expression of rhlR placing the las system in a cell-to-cell signaling hierarchy above the rhl system (Figure 4). Moreover, 3-oxo-C12-HSL can bind to RhlR, blocking the binding of C4-HSL to its transcriptional activator rhlR. The las system therefore controls the rhl system at both a transcriptional and posttranslational level. Another yet unidentified regulatory mechanism directly influencing the expression of rhlRI has been suggested. Rhl system regulation of such important genes as rpoS could explain why multiple levels of controls are required for its tight regulation.
Cell-to-Cell Signaling: A Powerful System to Overcome Host Defenses during Acute Infection
Cell-to-cell signaling systems might enable P. aeruginosa to overcome host defense mechanisms. Isolated production of extracellular virulence factors by a small number of bacteria would probably lead to an efficient host response neutralizing these compounds. However, the coordinated expression of virulence genes by an entire bacterial population once a certain density has been reached might allow P. aeruginosa to secrete extracellular factors only when they can be produced at high enough levels to overcome host defenses. These factors could alter the precarious balance between host defenses and production of bacterial toxins, leading to invasion of blood vessels, dissemination, systemic inflammatory-response syndrome, and finally death. Even appropriate antibiotic therapies are often unable to stop this course; therefore, the process must be blocked early, before virulence gene expression can be coordinated.
Biofilms and Cell-to-Cell Signaling
The most striking feature of persistent P. aeruginosa infections in CF patients is the selection of mucoid mutants producing the exopolysaccharide alginate (a polymer of mannuronic and guluronic acid). These mutant bacteria grow inside a biofilm (defined as microcolonies surrounded by an exopolysaccharide) as a survival strategy because the surrounding matrix protects bacteria from phagocytes and complement activity. P. aeruginosa growing in an alginate "slime matrix" are resistant to antibiotics (e.g., aminoglycosides, ъ-lactam antibiotics, fluoroquinolones) and disinfectants. The exact nature of the increased resistance is unclear but has been attributed to slow growth, penetration barriers, ъ-lactamase production, and other factors. P. aeruginosa also produces other less well-defined biofilms essential in the colonization of indwelling devices such as catheters. Recently, the las cell-to-cell signaling system has been shown to be involved in the differentiation of P. aeruginosa biofilms. A mutant defective in the production of 3-oxo-C12-HSL formed an abnormal biofilm that, in contrast to the wild type biofilm, was sensitive to low concentrations of the detergent sodium dodecyl sulfate (SDS). Furthermore, the addition of 3-oxo-C12-HSL to the culture media restored production of a differentiated, SDS-resistant biofilm by the mutant. Whether the formation of an undifferentiated biofilm renders this mutant more sensitive to antibiotics is still unknown. Also unclear is whether 3-oxo-C12-HSL is required for the differentiation of alginate biofilms. The link between 3-oxo-C12-HSL and biofilm differentiation highlights the broad range of systems controlled by cell-to-cell signaling in P. aeruginosa.